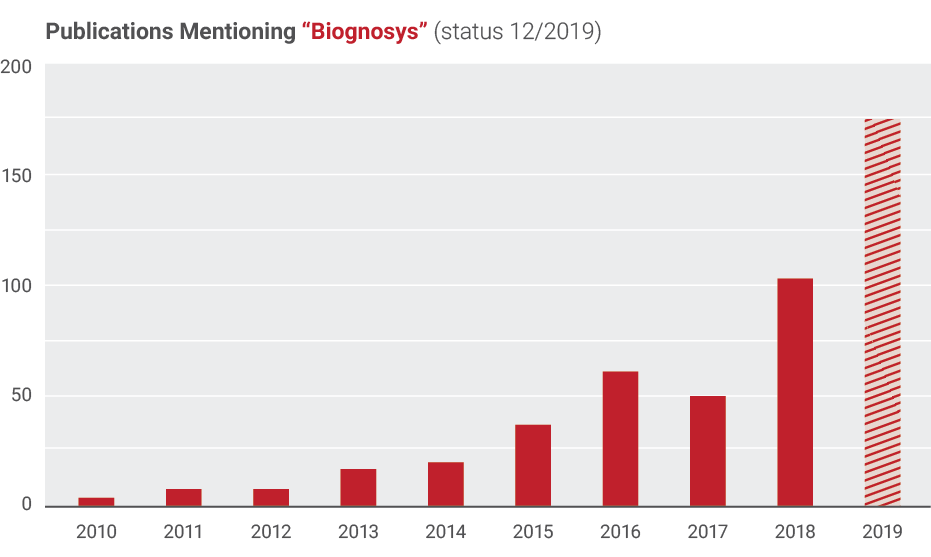
Publications by Biognosys and Collaborators
Bruderer R, Muntel J, Müller S, Bernhardt OM, Gandhi T, Cominetti O, Macron C, Carayol J, Rinner O, Astrup A, Saris WHM, Hager J, Valsesia A, Dayon L, and Reiter L. Molecular & Cellular Proteomics
We have established a robust capillary-flow DIA platform capable of measuring a large number of clinical samples in a fast and reproducible manner. The platform allowed the acquisition of 31 plasma proteomes per day, and a total of 1508 samples from the dietary intervention study DiOGenes measured on a single column. Our workflow revealed distinct biological reactions to weight loss and maintenance, and the comparison to independent studies showed the robustness for potential biomarker discovery.
Muntel J, Gandhi T, Verbeke L, Bernhardt OM, Treiber T, Bruderer R, and Reiter L. Molecular Omics
In this study we showed how improvements in chromatography and data analysis can push the proteome coverage in human tissue beyond 10,000 proteins, spanning six orders of magnitude in dynamic range. Such remarkable coverage was achieved using Hybrid Libraries, a novel approach that combines the depth of resource libraries with the high precision of directDIA. Thousands of these proteins could be linked to the hallmarks of testis cancer.
Muntel J, Kirkpatrick J, Bruderer R, Huang T, Vitek O, Ori A, and Reiter L. Journal of Proteome Research
Here we compare the performance of the two most popular quantitative workflows employed in discovery proteomics, label-free DIA, and isobaric-labeled TMT 10-plex. Workflows were tested on the same spike-in samples, with matched instrument time and reciprocal assessment in two facilities. Both workflows delivered high proteome coverage with low missing values at the protein level. While TMT performed slightly better in coverage and quantitative precision, DIA showed higher scalability and quantitative accuracy.
Huang T, Bruderer R, Muntel J, Xuan Y, Vitek O, and Reiter L. Molecular & Cellular Proteomics
In this publication, we report a novel statistical procedure that takes advantage of the two independent quantitative spaces available in DIA data, i.e., MS1 and MS2, for the detection of differentially abundant proteins. The use of these two levels improves precision and reduces the effect of the interferences. This new approach was applied to six sets of controlled mixtures, showing a consistent improvement in the ability to find differential changes at the protein level.
Publications Mentioning Biognosys or their Technology and Tools
Lundby A, Franciosa G, Emdal KB, Refsgaard JC, Gnosa SP, Bekker-Jensen DB, Secher A, Maurya SR, Paul I, Mendez BL, Kelstrup CD, Francavilla C, Kveiborg M, Montoya G, Jensen LJ, and Olsen J V. Cell
Researchers from the Olsen group developed an MS-based interaction proteomics approach to unravel the EGF-dependent phosphotyrosine signaling network in vivo. This approach is based on cutting-edge MS instrumentation and DIA, which together enabled the rapid and high-throughput screening of over a thousand phosphotyrosine sites. The authors showed how several cancer mutations near phosphotyrosine docking sites altered the assembly of protein-complexes and rewired signaling networks.
Amon S, Meier-Abt F, Gillet LCJ, Dimitrieva S, Theocharides APA, Manz MG, and Aebersold R. Molecular & Cellular Proteomics
Researchers from the Aebersold group presented a robust proteomic workflow for the quantitative profiling of minute amounts of sorted cells. Following isolation by fluorescence-activated cell sorting, the authors applied DIA to reproducibly quantify the proteome of hematopoietic stem and progenitor cells. The subpopulations of hematopoietic cells were analyzed in sets of 25,000, and on average over 5,800 protein groups were covered, representing a relevant technical and scientific advance.
Prosit: Proteome-wide Prediction of Peptide Tandem Mass Spectra by Deep Learning.
Gessulat S, Schmidt T, Zolg DP, Samaras P, Schnatbaum K, Zerweck J, Knaute T, Rechenberger J, Delanghe B, Huhmer A, Reimer U, Ehrlich H-C, Aiche S, Kuster B, and Wilhelm M. Nature Methods
Researchers at the Technical University of Munich trained a deep neural network, termed Prosit, to predict spectra for proteases, generate high-quality spectral libraries for DIA, and improve the analysis of metaproteomes. Prosit could learn and predict the retention time and fragment ion intensity of any peptide and has proven to exceed the quality of the experimental data. This tool can generate custom spectral libraries for any organism just from peptide sequence information.
Lysine Harvesting is an Antioxidant Strategy and Triggers Underground Polyamine Metabolism.
Olin-Sandoval V, Yu JSL, Miller-Fleming L, Alam MT, Kamrad S, Correia-Melo C, Haas R, Segal J, Peña Navarro DA, Herrera-Dominguez L, Méndez-Lucio O, Vowinckel J, Mülleder M, and Ralser M. Nature
Lipid Signaling Drives Proteolytic Rewiring of Mitochondria by YME1L.
MacVicar T, Ohba Y, Nolte H, Mayer FC, Tatsuta T, Sprenger H-G, Lindner B, Zhao Y, Li J, Bruns C, Krüger M, Habich M, Riemer J, Schwarzer R, Pasparakis M, Henschke S, Brüning JC, Zamboni N, and Langer T. Nature
Abundance of Bacterial Type VI Secretion System Components Measured by Targeted Proteomics.
Lin L, Lezan E, Schmidt A, and Basler M. Nature Communications
Mehnert M, Li W, Wu C, Salovska B, and Liu Y. Proteomics
Ye Z, Mao Y, Clausen H, and Vakhrushev SY. Nature Methods
Softic S, Meyer JG, Wang G-X, Gupta MK, Batista TM, Lauritzen HPMM, Fujisaka S, Serra D, Herrero L, Willoughby J, Fitzgerald K, Ilkayeva O, Newgard CB, Gibson BW, Schilling B, Cohen DE, and Kahn CR. Cell Metabolism
Fabre B, Korona D, Lees JG, Lazar I, Livneh I, Brunet M, Orengo CA, Russell S, and Lilley KS. Journal of Proteome Research
Metastatic-Niche Labeling Reveals Parenchymal Cells with Stem Features.
Ombrato L, Nolan E, Kurelac I, Mavousian A, Bridgeman VL, Heinze I, Chakravarty P, Horswell S, Gonzalez-Gualda E, Matacchione G, Weston A, Kirkpatrick J, Husain E, Speirs V, Collinson L, Ori A, Lee J-H, and Malanchi I. Nature
Systematic Comparison of Strategies for the Enrichment of Lysosomes by Data Independent Acquisition.
Singh J, Kaade E, Muntel J, Bruderer R, Reiter L, Thelen M, and Winter D. Journal of Proteome Research
Niu L, Geyer PE, Wewer Albrechtsen NJ, Gluud LL, Santos A, Doll S, Treit P V, Holst JJ, Knop FK, Vilsbøll T, Junker A, Sachs S, Stemmer K, Müller TD, Tschöp MH, Hofmann SM, and Mann M. Molecular Systems Biology